- PLAZA, a resource for plant comparative genomics. Proost et al., 2009 - Van Bel et al., 2012, 2018, 2022
|
 |
- PLAZA Diatoms, a resource for diatom comparative genomics. Vandepoele et al., 2013
|
|
- pico-PLAZA, a resource for algal comparative genomics. Osuna-Cruz et al., 2020
|
|
- TRAPID: an efficient online tool for the functional and comparative analysis of de novo RNA-Seq transcriptomes. Van Bel et al., 2013; Bucchini et al., 2021
|
 |
- Integrative inference of transcriptional networks in Arabidopsis yields novel ROS signalling regulators. De Clercq, Van de Velde et al., 2021. Nature Plants - Supplementary iGRN website
|
|
- Enhanced Maps of Transcription Factor Binding Sites Improve Regulatory Networks Learned from Accessible Chromatin Data. Kulkarni S, Jones M & Vandepoele K. Plant Physiology 2019 - Supplementary website
|
|
- A functional and evolutionary perspective on transcription factor binding in Arabidopsis thaliana. Heyndrickx KS, Van de Velde J, Wang C, Weigel D & Vandepoele K. Plant Cell 2014 - Supplementary website
|
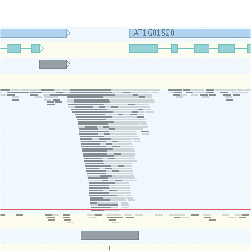 |
- Inference of transcriptional networks in Arabidopsis thaliana through conserved non-coding sequence analysis. Van de Velde J, Heyndrickx KS & Vandepoele K. Plant Cell 2014 - Supplementary website.

|
 |
- Systematic identification of functional plant modules through the integration of complementary data sources. Heyndrickx KS & Vandepoele K. Plant Physiology 2012 - Supplementary website
|
|
| For Tools & Supplementary data from papers before 2005, please check here.
|
|









