| IGR sequence: | igr1621#chr1#x1=6694921#l=2247 |
|---|---|
| Mature miRNA sequence: | GGUAGCCAAGGAUGACUUGCCU |
| encoded miRNA: | MIR43 |
| Precursor location: | 533 - 637 (negative strand) |
| precursor length: | 105 (37 basepairs) |
| MIR position: | 1 - 22 (616 - 637) |
| MIR length: | 22 (19 paired bases) |
| miRNA location TIGR v3: | chr1:6695453<6695557 |
| miRNA location TIGR v5: | chr1:6695453<6695557 |
| Folding energy: | -47.00 |
| BLAST hit against RFAM Atha miRNAs: | none |
| Belongs to miRNAs-targets cluster: | cluster006 |
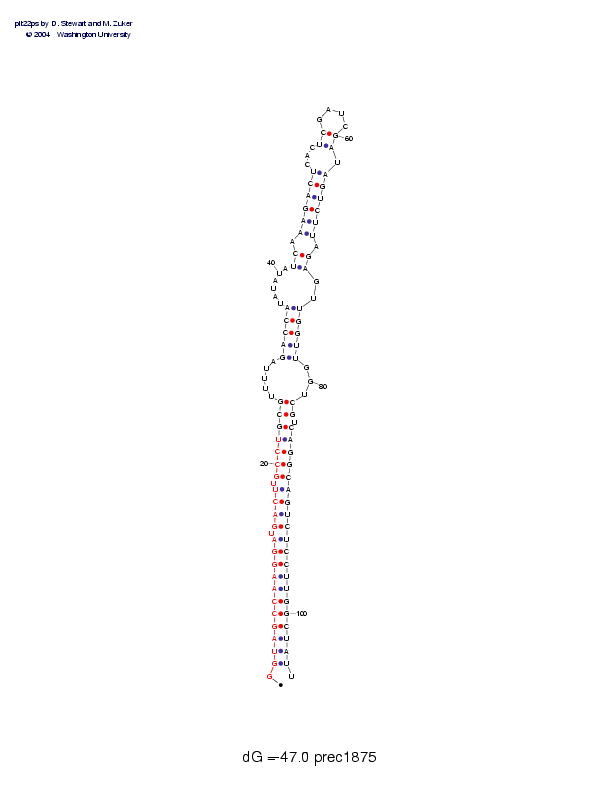
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
********** ********** **
GGUAGCCAAG GAUGACUUGC CUGCGUUUUA GACCAUAUAU AUCAAAGACU CACUCGAUCG 60
:((((((((( ((-((((-(( (((((----- (((((----- -((-(((((( ---((____)
AUAGUCUUAG AGUUGGUUGG UCGUCAGGCA GUCUCCUUGG CUAUU 105
)-))))))-) )--)))))-- -))-)))))) )))))))))) )))):
Postscript file CT file PNG file fasta file Stockholm file
Targets:
| Nr. | Gene | Description | mismatches | miRNAs |
|---|---|---|---|---|
| 1 | At1g17590.1 | 68408.m01959 transcription factor -related | 3 | 10 |
| 2 | At1g54160.1 | 68408.m05664 CCAAT-binding factor B subunit -related | 3 | 8 |
| 3 | At1g72830.1 | 68408.m07728 CCAAT-binding factor B subunit homolog -related | 3 | 8 |
Rice homologs
- gnl|BL_ORD_ID|2154_158028-158181_1-22
- gnl|BL_ORD_ID|2154_158030+158181_131-152
- gnl|BL_ORD_ID|323_84895+85046_131-152
- gnl|BL_ORD_ID|323_84893-85046_1-22