| IGR sequence: | igr2959#chr2#x1=14390339#l=3764 |
|---|---|
| Mature miRNA sequence: | UCUGCCAAAGGAGAUUUGCCC |
| encoded miRNA: | MIR27b |
| Precursor location: | 3137 - 3230 (positive strand) |
| precursor length: | 94 (37 basepairs) |
| MIR position: | 74 - 94 (3210 - 3230) |
| MIR length: | 21 (19 paired bases) |
| miRNA location TIGR v3: | chr2:14393475>14393568 |
| miRNA location TIGR v5: | chr2:14452088>14452181 |
| Folding energy: | -44.40 |
| BLAST hit against RFAM Atha miRNAs: | none |
| Belongs to miRNAs-targets cluster: | cluster020 |
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
GGGCAAGAUC ACCAUUGGCA GAGAUCUAUU ACUUCAUUCU UGCAUCAUAU GCAUAAAUGU 60
(((((-(((( -((-(((((( ((((-((((( ((--((((-- (((((___)) )))--))))-
******* ********** ****
UUGUGGUGAG CUCUCUGCCA AAGGAGAUUU GCCC 94
--)))))-)) -))))))))) )-))-))))) ))))
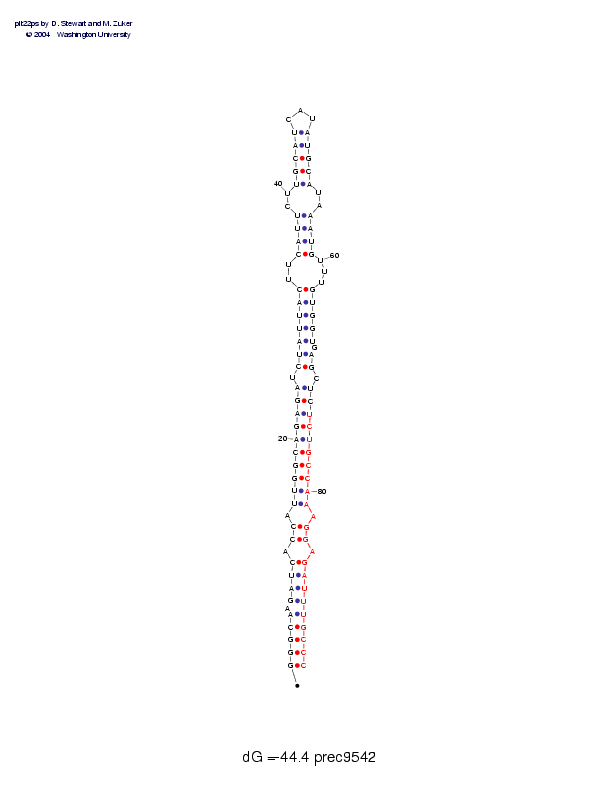
Postscript file CT file PNG file fasta file Stockholm file
Targets:
| Nr. | Gene | Description | mismatches | miRNAs |
|---|---|---|---|---|
| 1 | At2g33770.1 | 68409.m03744 ubiquitin-conjugating enzyme family | 2 | 5 |
| 2 | At2g33770.1 | 68409.m03744 ubiquitin-conjugating enzyme family | 2 | 5 |
Rice homologs
- gnl|BL_ORD_ID|302_10010-10104_1-21
- gnl|BL_ORD_ID|302_10010+10104_75-95
- gnl|BL_ORD_ID|289_138589-138683_1-21
- gnl|BL_ORD_ID|289_138589+138683_75-95