| IGR sequence: | igr4121#chr2#x1=19115407#l=13764 |
|---|---|
| Mature miRNA sequence: | UCGGACCAGGCUUCAUUCCC |
| encoded miRNA: | MIR5 |
| Precursor location: | 9178 - 9311 (positive strand) |
| precursor length: | 134 (48 basepairs) |
| MIR position: | 115 - 134 (9292 - 9311) |
| MIR length: | 20 (15 paired bases) |
| miRNA location TIGR v3: | chr2:19124584>19124717 |
| miRNA location TIGR v5: | chr2:19183197>19183330 |
| Folding energy: | -53.10 |
| BLAST hit against RFAM Atha miRNAs: | ath-MIR166a |
| Belongs to miRNAs-targets cluster: | cluster027 |
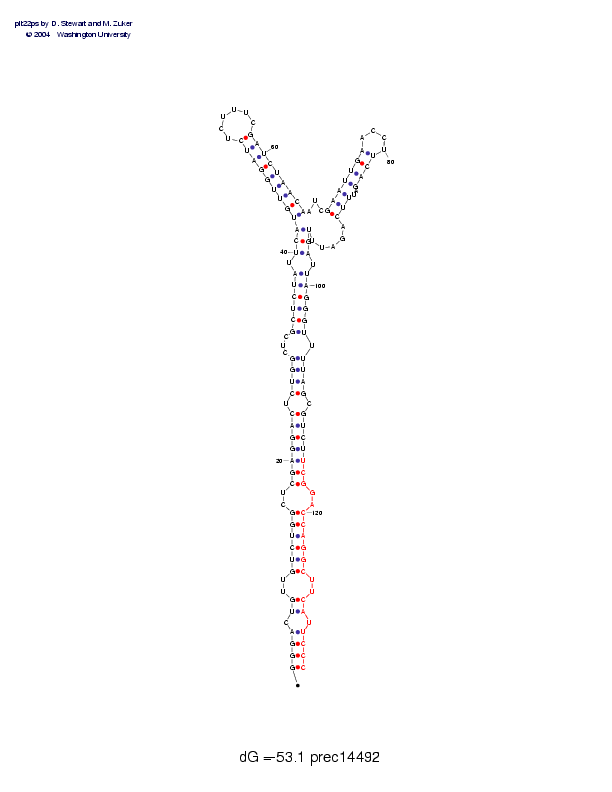
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
GGGACUGUUG UCUGGCUCGA GGACUCUGGC UCGCUCUAUU CAUGUUGGAU CUCUUUCGAU 60
((((-((--( (((((--((( ((((-((((- --((((((-( ((<<<<<<<< <______>>>
******
CUAACAAUCG AAUUGAACCU UCAGAUUUCA GAUUUGAUUA GGGUUUUAGC GUCUUCGGAC 120
>>>>>>,,,< <<<<<<____ >>>>-->>>, ,,,,)))-)) ))))-))))- )))))))--)
********** ****
CAGGCUUCAU UCCC 134
)))))--))- ))))
Postscript file CT file PNG file fasta file Stockholm file
Targets:
| Nr. | Gene | Description | mismatches | miRNAs |
|---|---|---|---|---|
| 1 | At1g52150.1 | 68408.m08708 HD-Zip transcription factor | 2 | 2 |
| 2 | At1g52150.2 | 68408.m08709 HD-Zip transcription factor | 2 | 2 |
Rice homologs
- gnl|BL_ORD_ID|1999_118134+118263_111-130
- gnl|BL_ORD_ID|1300_66425+66554_111-130