| IGR sequence: | igr2953#chr2#x1=14363467#l=4130 |
|---|---|
| Mature miRNA sequence: | UCCGUUUCACAAUAUAAGUU |
| encoded miRNA: | MIR73w |
| Precursor location: | 330 - 536 (positive strand) |
| precursor length: | 207 (55 basepairs) |
| MIR position: | 1 - 20 (330 - 349) |
| MIR length: | 20 (18 paired bases) |
| miRNA location TIGR v3: | chr2:14363796>14364002 |
| miRNA location TIGR v5: | chr2:14422409>14422615 |
| Folding energy: | -35.97 |
| BLAST hit against RFAM Atha miRNAs: | none |
| Belongs to miRNAs-targets cluster: | none |
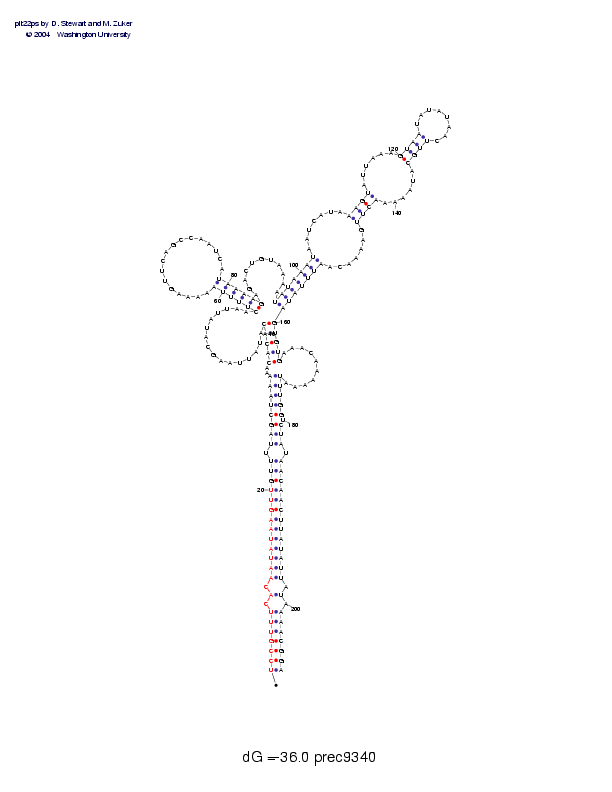
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
********** **********
UCCGUUUCAC AAUAUAAGUU GUUUUAGCUA AAAACACACA UAUUAAGCAU AUUAACUUUU 60
(((((((-(- (((((((((( (((-(((((( ((--(((((, ,,,,,,,,,, ,,,,,<<<<<
AAAAAGUUCA GCCAAUCAUA AAAGAGACUG UAAAAUAUAA AUAAUCAUAA AGUAUUAAAG 120
<_________ ________>> >>>>,,,,,, ,,,,,<<<<< <<-------< <<<------<
UAAUAUAUAA CUUGCAUAAA AACUUGAAAA CAAUUUAUAG UGUGAAACAA AAAAUUUGGU 180
<<<_______ _>>>>----- ->>>>----- -->>>>>>>) ))))------ ----)))))-
CUAUAACAAC UUAUAUUAUA AAACGGA 207
)))-)))))) )))))))-)- )))))))
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|3435_76530+76667_1-20
- gnl|BL_ORD_ID|3435_76530-76667_119-138