| IGR sequence: | igr2835#chr5#x1=14105322#l=9222 |
|---|---|
| Mature miRNA sequence: | CUUAUAUUGUAAAACGGAGG |
| encoded miRNA: | MIR51k |
| Precursor location: | 3188 - 3383 (positive strand) |
| precursor length: | 196 (70 basepairs) |
| MIR position: | 177 - 196 (3364 - 3383) |
| MIR length: | 20 (19 paired bases) |
| miRNA location TIGR v3: | chr5:14049209>14049404 chr5:14108509>14108704 |
| miRNA location TIGR v5: | chr5:14458267>14458462 chr5:14517567>14517762 |
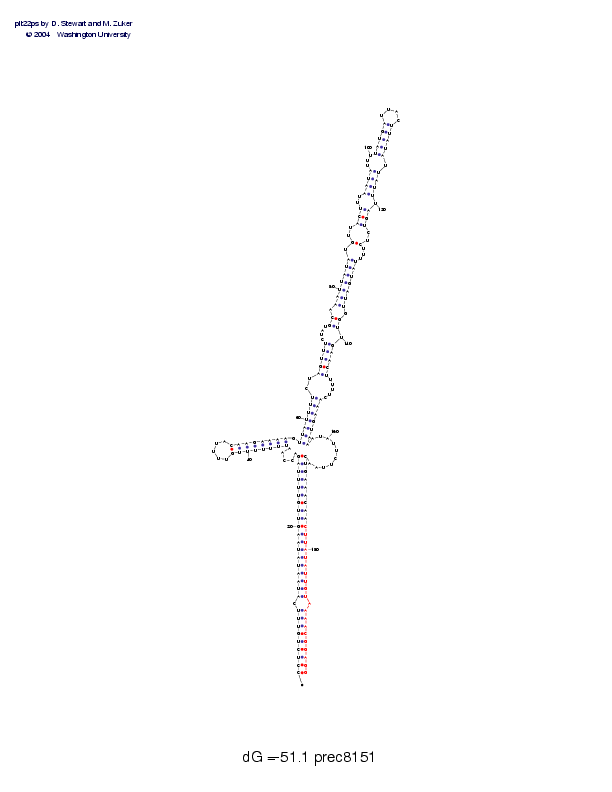
| Folding energy: | -51.10 |
| BLAST hit against RFAM Atha miRNAs: | none |
| Belongs to miRNAs-targets cluster: | none |
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
CCUCUGUUUC AUAAUAUAAG UUGUUUUAGA CCAAUUUUUU UGUUUUACAA GAAAAGUUAU 60
(((((((((- (((((((((( (((((((((, ,,,,<<<<<< <<_____>>> >>>>>,<<<<
UUUCUAGUUU CUAUGCAAAU UAUAUGUUAC UUUAAUAUUU UAUGAUUACU UAUAUUAUUU 120
<<<--<<<<< ----<<-<<< <<<<-<--<< <--<<<<--- <<<<<____> >>>>->>>>-
****
AGUCUCUUUU AUGAUUGGUU GAACUUUUUC AAAGUAAAUA UUCUUAACUG AAACAACUUA 180
>>>-->---> >>>>>>->>- >>>>>----- >>>>>>>,,, ,,,,,,,))) ))))))))))
********** ******
UAUUGUAAAA CGGAGG 196
))))))-))) ))))))
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|3371_67633+67752_101-120
- gnl|BL_ORD_ID|3371_67633-67752_1-20
- gnl|BL_ORD_ID|2045_114644-114763_1-20
- gnl|BL_ORD_ID|2045_114644+114763_101-120
- gnl|BL_ORD_ID|3458_114689-114785_1-20
- gnl|BL_ORD_ID|3458_114689+114785_78-97
- gnl|BL_ORD_ID|2537_132225+132321_78-97
- gnl|BL_ORD_ID|2537_132225-132321_1-20