| IGR sequence: | igr2102#chr2#x1=10621377#l=6616 |
|---|---|
| Mature miRNA sequence: | UGUGUGCUCACUCUCUUCUGUCAG |
| encoded miRNA: | MIR63a |
| Precursor location: | 3561 - 3644 (positive strand) |
| precursor length: | 84 (34 basepairs) |
| MIR position: | 61 - 84 (3621 - 3644) |
| MIR length: | 24 (24 paired bases) |
| miRNA location TIGR v3: | chr2:10624937>10625020 |
| miRNA location TIGR v5: | chr2:10683550>10683633 |
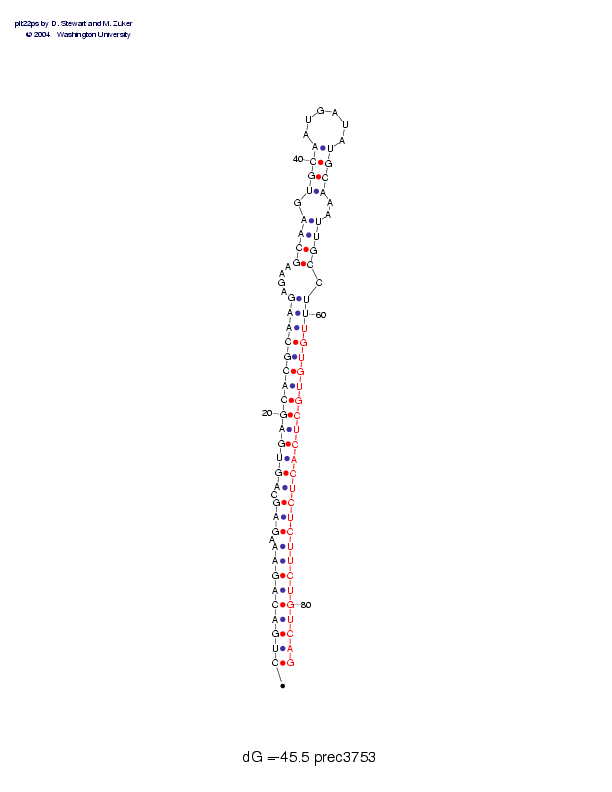
| Folding energy: | -45.50 |
| BLAST hit against RFAM Atha miRNAs: | ath-MIR156a |
| Belongs to miRNAs-targets cluster: | none |
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
CUGACAGAAA GAGCAGUGAG CACGCAAGAG AAGCAAGUGC AAUGAUAUGC AAAUUGCCUU 60
(((((((((- (((-(((((( ((((((((-- --((((-((( (______))) )--))))-))
********** ********** ****
UGUGUGCUCA CUCUCUUCUG UCAG 84
)))))))))) )))))))))) ))))
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|2024_146440+146551_89-112
- gnl|BL_ORD_ID|2024_146440-146551_1-24
- gnl|BL_ORD_ID|2021_81254-81365_1-24
- gnl|BL_ORD_ID|2021_81254+81365_89-112
- gnl|BL_ORD_ID|490_72534-72621_1-24
- gnl|BL_ORD_ID|490_72538-72621_1-24
- gnl|BL_ORD_ID|490_72534+72621_65-88
- gnl|BL_ORD_ID|490_72532+72621_67-90
- gnl|BL_ORD_ID|490_72149-72241_1-24
- gnl|BL_ORD_ID|490_72149+72241_70-93
- gnl|BL_ORD_ID|490_72146+72241_73-96
- gnl|BL_ORD_ID|457_25563-25650_1-24
- gnl|BL_ORD_ID|457_25567-25650_1-24
- gnl|BL_ORD_ID|457_25563+25650_65-88
- gnl|BL_ORD_ID|457_25561+25650_67-90
- gnl|BL_ORD_ID|457_25178-25270_1-24
- gnl|BL_ORD_ID|457_25178+25270_70-93
- gnl|BL_ORD_ID|457_25175+25270_73-96