| IGR sequence: | igr2044#chr5#x1=9109684#l=5223 |
|---|---|
| Mature miRNA sequence: | UGUGUGCUCACUCUCUUCUGUCA |
| encoded miRNA: | MIR17e |
| Precursor location: | 4190 - 4274 (negative strand) |
| precursor length: | 85 (30 basepairs) |
| MIR position: | 63 - 85 (4190 - 4212) |
| MIR length: | 23 (19 paired bases) |
| miRNA location TIGR v3: | chr5:9113873<9113957 |
| miRNA location TIGR v5: | chr5:9136129<9136213 |
| Folding energy: | -38.00 |
| BLAST hit against RFAM Atha miRNAs: | ath-MIR156f |
| Belongs to miRNAs-targets cluster: | none |
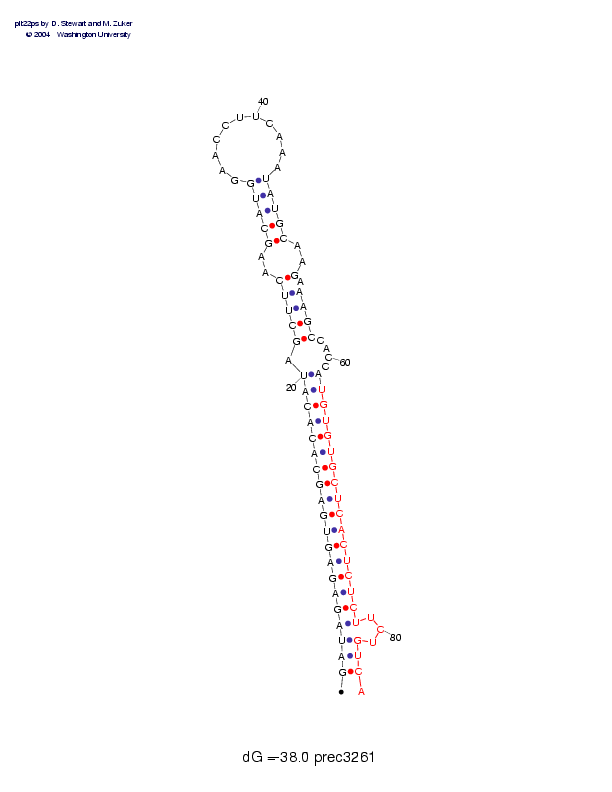
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
GAUAGAGAGU GAGCACACAU AGCUUCAAGC AUGGAACCUU CAAAUAUGCA AGAAAGCCAC 60
(((((((((( (((((((((( -(((((--(( (((_______ ____)))))- -)-))))---
******** ********** *****
CAUGUGUGCU CACUCUCUUC UGUCA 85
-))))))))) ))))))))-- -))):
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|2024_146528-146637_88-110
- gnl|BL_ORD_ID|2024_146528+146637_1-23
- gnl|BL_ORD_ID|2021_81343+81452_1-23
- gnl|BL_ORD_ID|2021_81343-81452_88-110