| IGR sequence: | igr1656#chr3#x1=6684108#l=3037 |
|---|---|
| Mature miRNA sequence: | UCAUUACUUCCAAAUACAGA |
| encoded miRNA: | MIR42 |
| Precursor location: | 1861 - 2210 (positive strand) |
| precursor length: | 350 (115 basepairs) |
| MIR position: | 331 - 350 (2191 - 2210) |
| MIR length: | 20 (20 paired bases) |
| miRNA location TIGR v3: | chr3:6685968>6686317 |
| miRNA location TIGR v5: | chr3:6685968>6686317 |
| Folding energy: | -128.65 |
| BLAST hit against RFAM Atha miRNAs: | none |
| Belongs to miRNAs-targets cluster: | none |
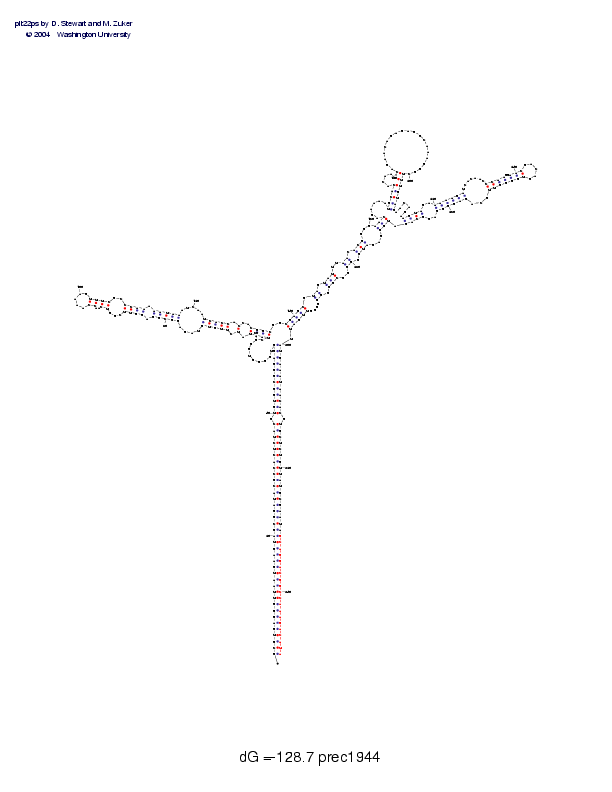
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
UCUGUAUUUG GAAGUAAUGA UCUUAGACUC CACGCGACUG UGUUCUUUAU UGUUUUGUUU 60
(((((((((( (((((((((( (((((((((( ((((((((-( (((((((((( (,,,,,,,,,
CGAAGAGCGG UGUGAUAUUC UUAUUGGCUU GCAACCAAAA UGGGCUCCCA AAAAGAAAGC 120
,<<<<-<-<< <<<----<<< <<-<<<<--- <<--<<____ _>>>>-->>> >->>>>>---
AAGCACCACC ACUUCUUCUA UCUGAUGAGG GAACACUUUU AACAUAAAAG UCACACAACC 180
-->>>>>->- ->>>>,,<<< <<-<<-<<<- -<<-<<<--- <<<,,,,,,[ [[[[----[[
AUAACAUAUA UUCAAAUUUC GGGUGAUAAC AUAAUGAUAA AAAUAGCACA ACCCAAUUAA 240
__________ __________ ]]]]]]],,, ,,,[[[[--[ [[[[[----- --[[[[--[[
CUACAUGUUU UGGUCAUAUU UUAUUCAUAG UUUAUAGUUU UUUCUUUUUU UUUGGUAGGA 300
[_____]]]] ]]]---]]]] ]]--]]]],> >>--->>>-> >-->>>->>- --->>>>>,)
********** **********
GUAAAGAACA CUGUCGCGUG GAGUCUAAGA UCAUUACUUC CAAAUACAGA 350
)))))))))) )-)))))))) )))))))))) )))))))))) ))))))))))
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|1622_98566+98831_247-266
- gnl|BL_ORD_ID|1480_156180+156445_247-266