| IGR sequence: | igr3768#chr5#x1=18615936#l=2646 |
|---|---|
| Mature miRNA sequence: | GUGGCAUACAGGGAGCCAGGCA |
| encoded miRNA: | MIR75i |
| Precursor location: | 1332 - 1412 (positive strand) |
| precursor length: | 81 (30 basepairs) |
| MIR position: | 60 - 81 (1391 - 1412) |
| MIR length: | 22 (19 paired bases) |
| miRNA location TIGR v3: | chr5:18617267>18617347 |
| miRNA location TIGR v5: | chr5:19026325>19026405 |
| Folding energy: | -42.30 |
| BLAST hit against RFAM Atha miRNAs: | ath-MIR160c |
| Belongs to miRNAs-targets cluster: | none |
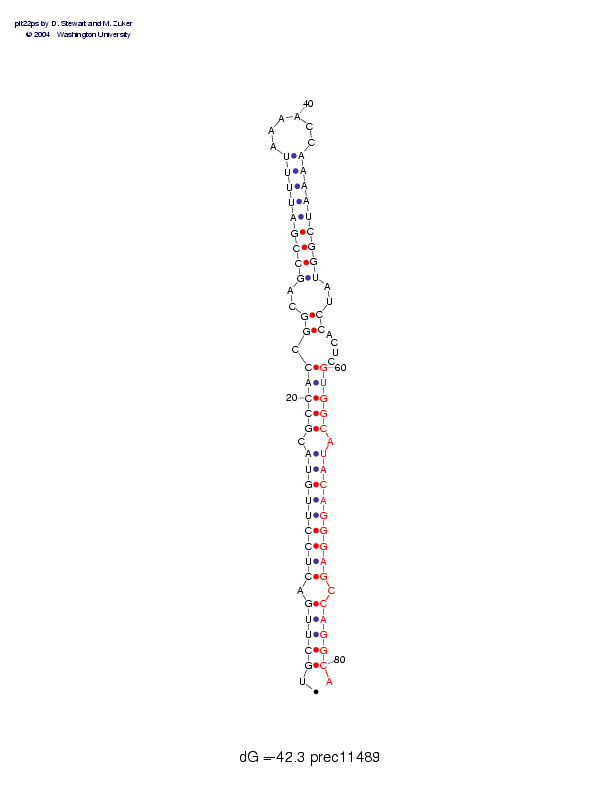
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
*
UGCUUGACUC CUUGUACGCC ACCGGCAGCC GAUUUUAAAA CCAAAAUCGG UAUCCACUCG 60
:(((((-((( ((((((-((( ((-((--((( ((((((____ __)))))))) )--))----)
********** ********** *
UGGCAUACAG GGAGCCAGGC A 81
))))-))))) ))))-))))) :
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|1495_131987+132068_61-82
- gnl|BL_ORD_ID|1495_131987-132068_1-22
- gnl|BL_ORD_ID|1461_56381+56462_61-82
- gnl|BL_ORD_ID|1461_56381-56462_1-22
- gnl|BL_ORD_ID|1294_49693-49779_1-22
- gnl|BL_ORD_ID|727_84855-84941_1-22