| IGR sequence: | igr3351#chr2#x1=16006446#l=6289 |
|---|---|
| Mature miRNA sequence: | GGCGCUAUCCCUCCUGAGCUU |
| encoded miRNA: | MIR26a |
| Precursor location: | 3991 - 4061 (negative strand) |
| precursor length: | 71 (23 basepairs) |
| MIR position: | 51 - 71 (3991 - 4011) |
| MIR length: | 21 (17 paired bases) |
| miRNA location TIGR v3: | chr2:16010436<16010506 |
| miRNA location TIGR v5: | chr2:16069049<16069119 |
| Folding energy: | -33.70 |
| BLAST hit against RFAM Atha miRNAs: | none |
| Belongs to miRNAs-targets cluster: | none |
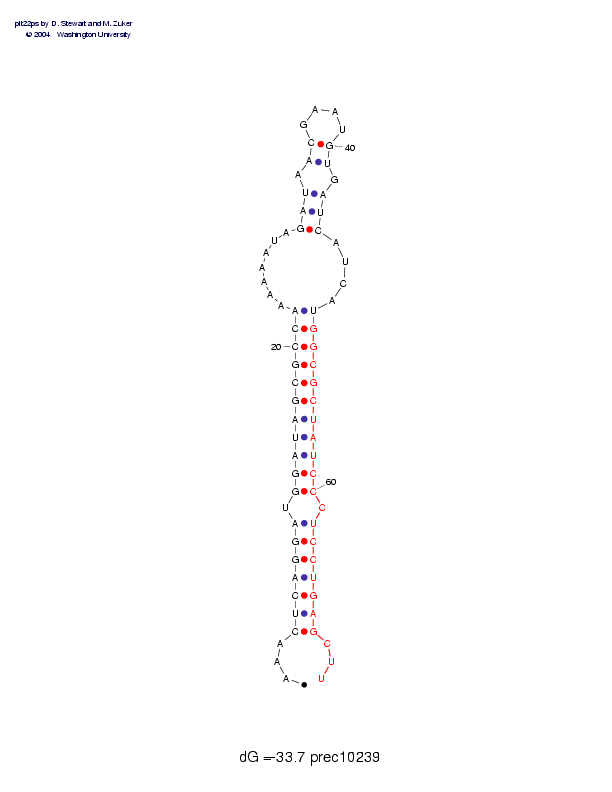
Sequence and secondary structure:
Presumed mature miRNA positions are indicated by *'s.
**********
AAACUCAGGA UGGAUAGCGC CAAAAAAUAG AUAACGAAUG UGAUCAUCAU GGCGCUAUCC 60
:::((((((( -((((((((( ((-------( ((-((____) )-)))----) ))))))))))
********** *
CUCCUGAGCU U 71
-))))))):: :
Postscript file CT file PNG file fasta file Stockholm file
Targets:
No Targets found for this miRNA
Rice homologs
- gnl|BL_ORD_ID|3598_86365+86419_1-21
- gnl|BL_ORD_ID|3598_86365-86420_36-56