Ectocarpus siliculosus
Taxonomy: Eukaryota; stramenopiles; PX clade; Phaeophyceae; Ectocarpales; Ectocarpaceae; Ectocarpus
Introduction
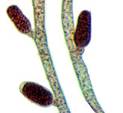
The brown algae are of interest for a number of reasons ranging from their economic importance as a biological resource to their phylogenetic position within the eukaryotes. Particular points of interest, for example, include the independent evolution of complex multicellularity within this lineage, the secondary endosymbiosis event that produced chloroplasts and the production of unique biomolecules.
In a recent study, the algal Genetics group in Roscoff (France) proposed Ectocarpus siliculosus as a model organism for the application of genetic and genomic approaches to the brown algae (Peters et al., J. Phycol. 2004). This proposition was based on several features of Ectocarpus including :
- its small size (several centimetres)
- the fact that its life cycle can be completed in culture in less than 3 months
- genetic analysis (crosses, complementation) is possible
- large scale mutant screens are possible (several hundred young plants can be screened in a single petri dish for example)
- it is highly fertile
- it is infected by an integrating DNA virus (potentially providing a tool for insertional mutagenesis)
- it has a relatively small genome size of 214 Mbp
Ectocarpus Genome Project
 In June 2004 an international consortium of 34 laboratories, coordinated in Roscoff, submitted a whole genome sequencing project to the French sequencing centre Genoscope. The project was accepted in September 2004 and sequencing of a 10x coverage of the 214 Mbp genome is currently in progress. The project will also include 150,000 reads on cDNA libraries constructed from mRNA corresponding to a range of developmental stages and treatments. In June 2004 an international consortium of 34 laboratories, coordinated in Roscoff, submitted a whole genome sequencing project to the French sequencing centre Genoscope. The project was accepted in September 2004 and sequencing of a 10x coverage of the 214 Mbp genome is currently in progress. The project will also include 150,000 reads on cDNA libraries constructed from mRNA corresponding to a range of developmental stages and treatments.
Our involvement
The annotation of the Ectocarpus genome will be done in our group in collaboration with the other members of the Ectocarpus consortium. To produce automated annotations we will adapt the EuGene system to the specificities of the Ectocarpus genome in order to produce a high quality annotation. Full exploitation of the genome sequence will require the establishment of a strong bioinformatics and biological infrastructure and this process has been initiated within the context of the Ectocarpus Genome Consortium. To communicate the annotations to the consortium a bioinformatics portal will be installed to facilitate the expert manual annotations and to offer a variety of bioinformatics tools and services for the functional analysis of the Ectocarpus genome.
In collaboration with:
Publications
2011Cock, M., Colle, J., Sterck, L., Rouzé, P., Scornet, D., Anthouard, V., Artiguenave, F., Aury, J., Billiau, K., Bonnet, E., H.F. Bothwell, J., Brillet, L., Carre, W., Coelho, S.M., Corre, E., Da Silva, C., Jubin, C., Martens, C., Maumus, F., Miranda-Saavedra, D., Peters, S., Porcel, B., Quesneville, H., Boyen, C., Van de Peer, Y., Wincker, P. (2011)
Nature, nurture and the structure of macroalgal genomes. Eur. J. Phycol. 46,39-39.
2010Cock, M., Sterck, L., Rouzé, P., Scornet, D., Allen, A., Amoutzias, G., Anthouard, V., Artiguenave, F., Aury, J., Badger, J., Beszteri, B., Billiau, K., Bonnet, E., Bothwell, J., Bowler, C., Boyen, C., Brownlee, C., Carrano, C., Charrier, B., Youn Cho, G., Coelho, S.M., Colln, J., Corre, E., Da Silva, C., Delage, L., Delaroque, N., Dittami, S., Doulbeau, S., Elias, M., Farnham, G., Gachon, C., Gschloessl, B., Heesch, S., Jabbari, K., Jubin, C., Kawai, H., Kimura, K., Kloareg, B., Küpper, F.C., Lang, D., Le Bail, A., Leblanc, C., Lerouge, P., Lohr, M., Lopez, P., Martens, C., Maumus, F., Michel, G., Miranda-Saavedra, D., Morales, J., Moreau, H., Motomura, T., Nagasato, C., Napoli, C., Nelson, D., Nyvall-Collén, P., Peters, S., Pommier, C., Potin, P., Poulain, J., Quesneville, H., Read, B., Rensing, S.A., Ritter, A., Rousvoal, S., Samanta, M., Samson, G., Schroeder, D., Ségurens, B., Strittmatter, M., Tonon, T., Tregear, J., Valentin, K., von Dassow, P., Yamagishi, T., Van de Peer, Y., Wincker, P. (2010)
The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature 465(7298):617-21.
2009Dittami, S., Scornet, D., Petit, J., Ségurens, B., Da Silva, C., Corre, E., Dondrup, M., Glatting, K., Konig, R., Sterck, L., Rouzé, P., Van de Peer, Y., Cock, M., Boyen, C., Tonon, T. (2009)
Global expression analysis of the brown alga Ectocarpus siliculosus (Phaeophyceae) reveals large-scale reprogramming of the transcriptome in response to abiotic stress. Genome Biol. 10(6):R66.
|


Internal Links
|
 The brown algae are of interest for a number of reasons ranging from their economic importance as a biological resource to their phylogenetic position within the eukaryotes. Particular points of interest, for example, include the independent evolution of complex multicellularity within this lineage, the secondary endosymbiosis event that produced chloroplasts and the production of unique biomolecules.
The brown algae are of interest for a number of reasons ranging from their economic importance as a biological resource to their phylogenetic position within the eukaryotes. Particular points of interest, for example, include the independent evolution of complex multicellularity within this lineage, the secondary endosymbiosis event that produced chloroplasts and the production of unique biomolecules.

