A functional and evolutionary perspective on transcription factor binding in Arabidopsis thalianaKen S Heyndrickx*, Jan Van de Velde*, Congmao Wang, Detlef Weigel and Klaas Vandepoele*Contributed equallyCorresponding author: AbstractUnderstanding the mechanisms underlying gene regulation is paramount to comprehend the translation from genotype to phenotype. The two are connected by gene expression, and it is generally thought that variation in transcription factor (TF) function is an important determinant of phenotypic evolution. We analysed publicly available genome-wide ChIP experiments for 27 transcription factors (TFs) in Arabidopsis thaliana and constructed an experimental network containing 46,619 regulatory interactions and 15,188 target genes. Hub targets and Highly Occupied Target (HOT) regions were identified, which are enriched for genes involved in development, stimuli response, signaling and gene regulatory processes. We provide several lines of evidence that TF binding at plant HOT regions is functional, in contrast to animals, and not merely the result of accessible chromatin. HOT regions harbor specific DNA motifs, are enriched for differentially expressed genes, are often conserved across crucifers and dicots, even though they are not under higher levels of purifying selection than non-HOT regions. Distal bound regions are under purifying selection as well, and are enriched for a chromatin state showing regulation by the Polycomb repressive complex. Gene expression complexity is positively correlated with the total number of regulating TFs, revealing insights in the regulatory code for genes with different expression breadths. The integration of non-canonical and canonical DNA motif information yields new hypotheses on co-binding and tethering between specific TFs involved in flowering and light regulation. Reference: A functional and evolutionary perspective on transcription factor binding in Arabidopsis thaliana. Heyndrickx KS, Van de Velde J, Wang C, Weigel D, Vandepoele K. Plant Cell. 2014 Oct;26(10):3894-910. Supplementary Data
A quick overview of the different tracks and features in the genome browser are displayed below. An extended Genomeview manual can be found here. 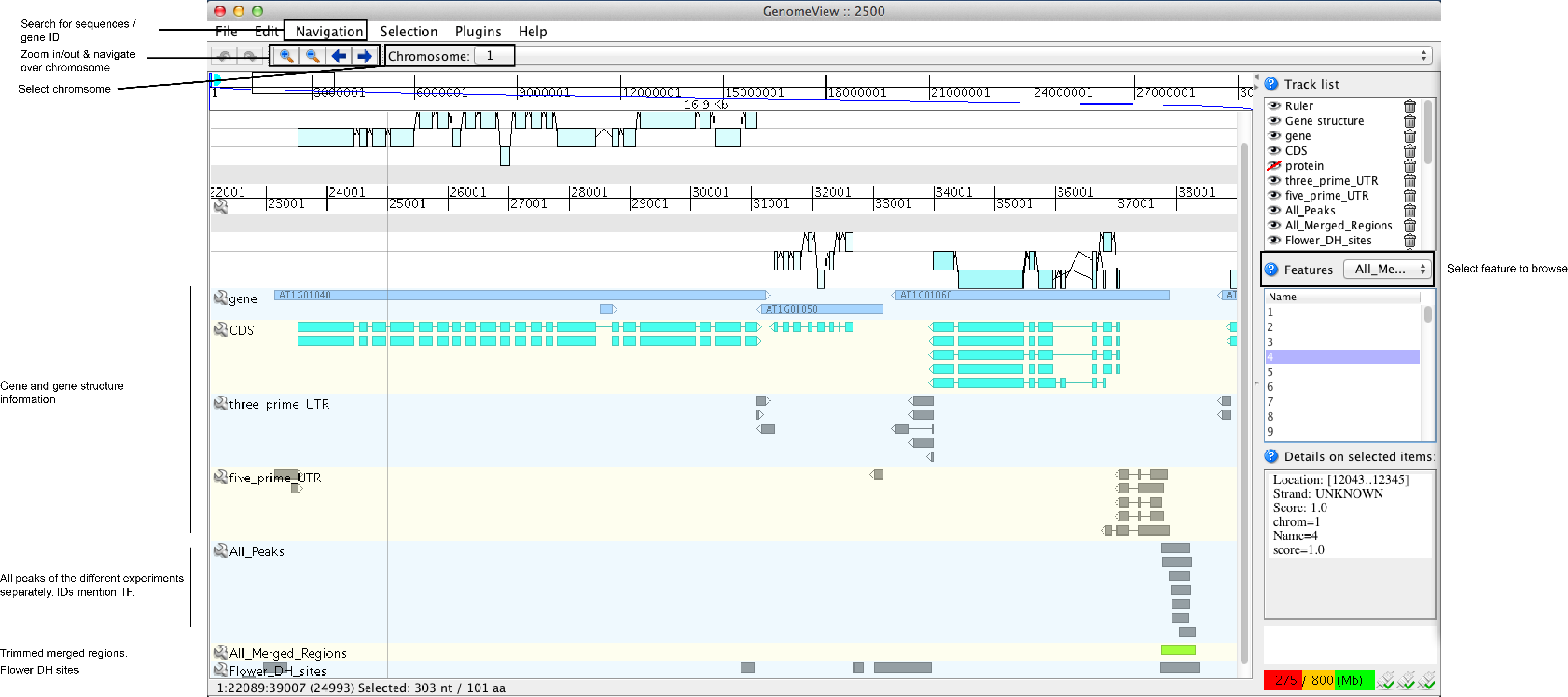
|