Motivation:
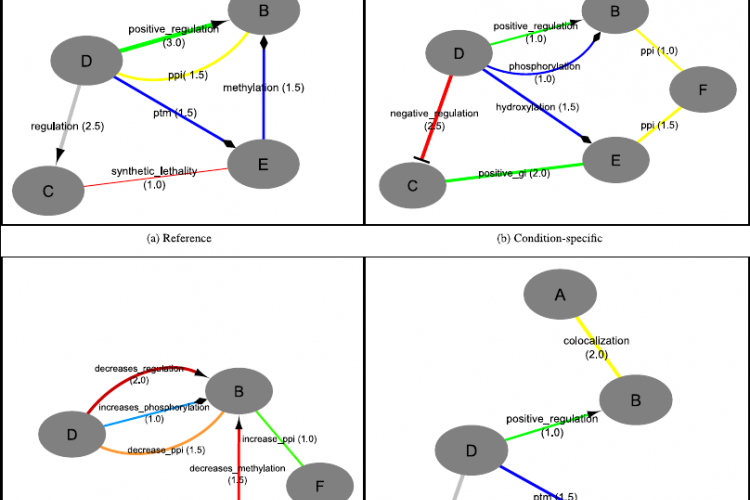
Differential networks have recently been introduced as a powerful way to study the dynamic rewiring capabilities of an interactome in response to changing environmental conditions or stimuli. Currently, such differential networks are generated and visualised using ad hoc methods, and are often limited to the analysis of only one condition-specific response or one interaction type at a time.
Results:
In this work we present an ontology-driven framework to infer and visualise an arbitrary set of condition-specific responses against a reference network. To this end, we have implemented algorithms that can process highly heterogeneous networks, accounting for both physical interactions and regulatory associations, symmetric and directed edges, edge weights and negation. We propose this integrative framework as a standardized methodology that allows a unified view on differential networks and promotes comparability between differential network studies. As an illustrative application, we demonstrate its usefulness on a plant abiotic stress study and experimentally confirmed a predicted regulator.
Van Landeghem, S., Van Parys, T., Dubois, M., Inzé, D., & Van de Peer, Y. (2016). Diffany: an ontology-driven framework to infer, visualise and analyse differential molecular networks. BMC BIOINFORMATICS, 17.