| |
Home |
| |
Manual |
| |
Download |
| |
History |
|
Latest update/bug fix: May 13,
2008 (see History) If you use ENIGMA in your research, please cite: Steven Maere, Patrick Van Dijck, Martin Kuiper (2008) Extracting expression modules from perturbational gene expression compendia. BMC Systems Biology 2:33 (PubMed, BMC) |
|
|
ENIGMA is a software tool to extract gene expression modules from perturbational microarray data, based on the use of combinatorial statistics and graph-based clustering. The modules are further characterized by incorporating other data types, e.g. GO annotation, protein interactions and transcription factor binding information, and by suggesting regulators that might have an effect on the expression of (some of) the genes in the module.
Current software version : ENIGMA 1.1
Current GO annotation version : Aug 29th 2007
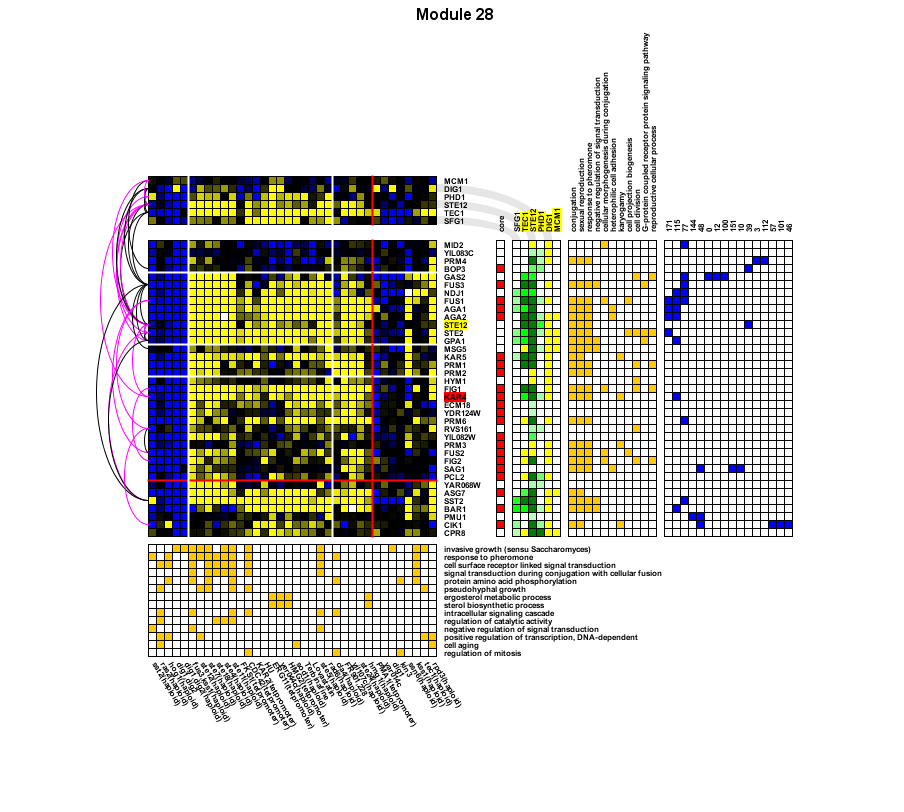
Below is a sample output figure for a mating-related module learned from the Rosetta dataset (Hughes et al (2000) Cell 102:109-126) |
|
|
|
|
|
Copyright (c) 2007 Flanders Interuniversitary Institute for Biotechnology (VIB) |